Research
Our research focuses on two overall questions:
Q1: How do normal, healthy cells evolve to become cancer cells?
Q2: How do we translate predicted cancer risk to patients effectively?
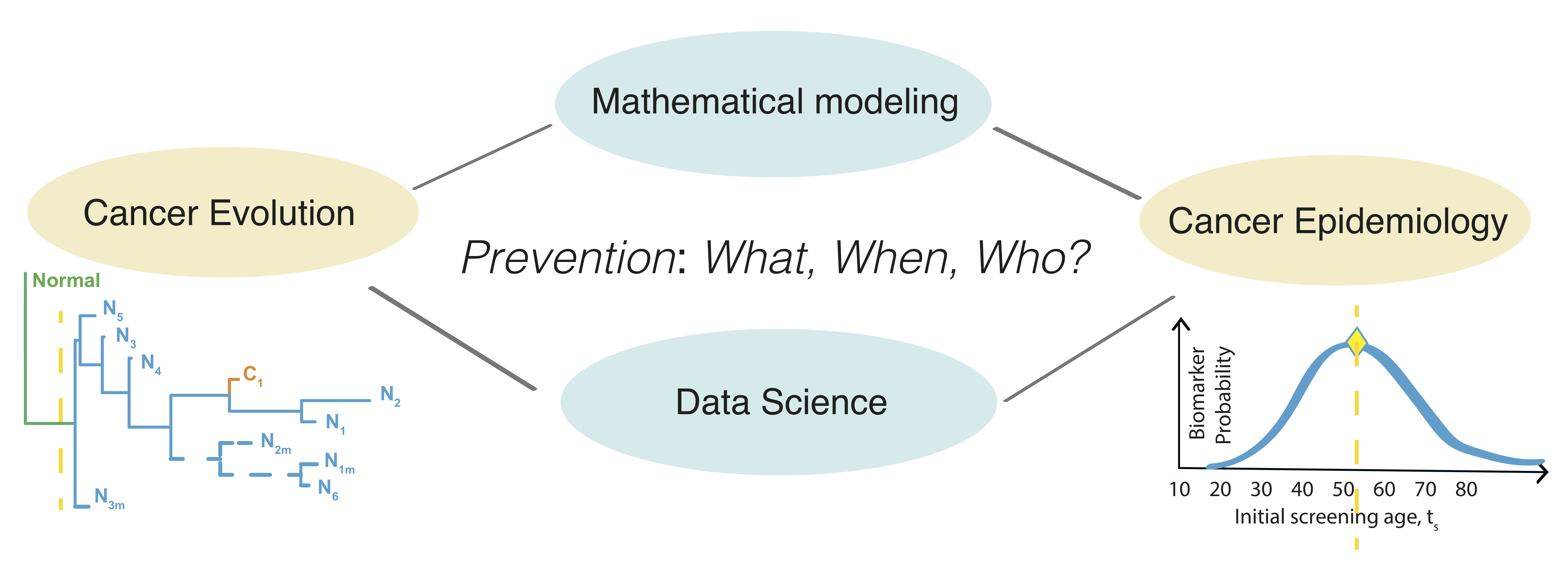
These two questions have traditionally been examined in isolation from one another. However, molecular and population data available are inextricably linked - the underlying dynamic process of cancer evolution manifests in the human body and measurements may be naturally reflected in incidence or in collected patient snapshots based on the timing of clinical recommmendations/study design, and interventions in practice.
We leverage these inherent connections by using mechanistic models, machine learning, and bioinformatics methods to integrate epidemiological, clinico-pathological and molecular cancer data in order to test and validate hypotheses. With a current focus on gastrointestinal cancers, we seek a coherent understanding of the carcinogenesis process so we may find and identify dangerous precursors (or diverse hotbeds of clones) in patients and forecast outcomes reliably.
The following are some examples of our collaborative research in GI cancer control:
What are the premalignant alterations that are predictive of future cancers?
Methods:
Genome sequencing, bioinformatics, phylogenetics, biostatistics, predictive modeling, longitudinal data analysis
Reference links with applications:
- Copy number alterations predict cancer in inflammatory bowel disease
- Efficacy of screening modalities for detecting high-risk dysplasia in Barrett’s esophagus
When are patients most at risk of premalignancy and would benefit from screening?
Methods:
Mechanistic modeling, stochastic processes, Bayesian inference of biological aging, cost-effectiveness analyses
Reference links with applications:
- Patient-specific molecular clocks for Barrett’s esophagus onset ages
- Optimal timing of screening and adaptive surveillance
- Estimating timescales of (pre)malignant growth from incidence curves
Who should we refer for screening and who then requires further interventions?
Methods:
Risk stratification, translational bioinformatics, evolutionary biomarkers, clinical decision support tools
Reference links with applications:
- UC-CaRE: a validated colitis-associated colorectal cancer risk prediction web-tool
- Modeling suggests Barrett’s esopohagus patients constitute most future esophageal adenocarcinomas
For further reading about our research, please see our section on “Cancer screening and early detection with modeling” as part of the 2019 Mathematical Oncology Roadmap.
